Citations
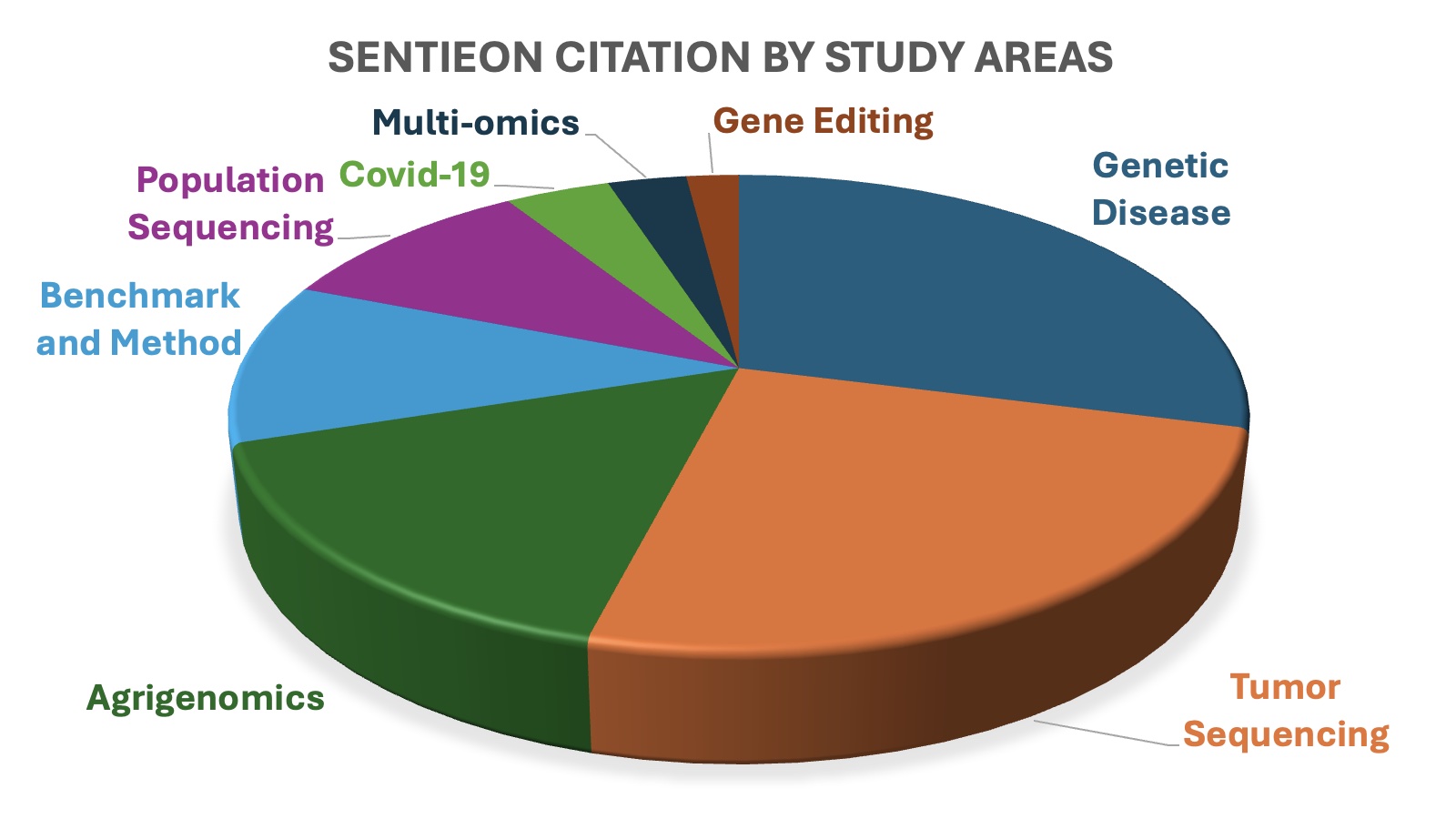
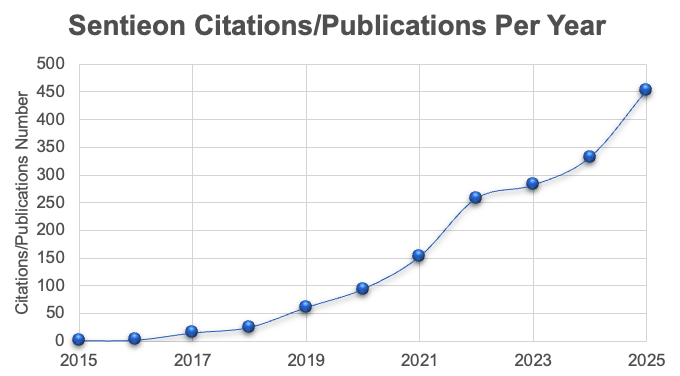
More than 1500 publications citing the Sentieon Software from multiple study areas
-
Genetic Disease
Clinical whole genome, exome, and panel sequencing. Selected Citation List
-
Tumor Sequencing
Accurately identify variants from fresh, frozen, and FFPE biopsies or blood samples. Selected Citation List
-
Agrigenomics
Analyze complex and polyploid plant and animal genomes. Selected Citation List
-
Population Sequencing
Cost-effective whole genome and exome sequencing joint genotyping of large cohorts. Selected Citation List
-
Multi-omics
Analyze genomic variants, gene expression, methylation and other datasets simultaneously. Selected Citation List
-
Gene Editing
Screen whole genome for off-target events and calculate editing efficiency. Selected Citation List
Sentieon Publications and Preprints
To Cite Sentieon® tools, please consider the corresponding pipeline publications listed below.
- [DNAscope for ONT Dataset - BioRxiv 2025] - Accelerated and High-Accuracy Variant Calling on Oxford Nanopore Technologies Sequencing Data with the Sentieon DNAscope LongRead and Hybrid Pipelines
- [DNAscope Hybrid - Frontiers in Bioinformatics 2025] - A Novel and Accelerated Method for Integrated Alignment and Variant Calling from Short and Long Reads
- [non-UMI ctDNA - BioRxiv 2024] - Optimizing Accuracy and Efficiency in Analyzing Non-UMI Liquid Biopsy Datasets Using the Sentieon ctDNA Pipeline
- [Gene Editing - JGG 2023] - CRISPR-detector: fast and accurate detection, visualization, and annotation of genome-wide mutations induced by genome editing events
- [DNAscope - BioRxiv 2022] - DNAscope: High accuracy small variant calling using machine learning
- [DNAscope LongRead - BioRxiv 2022] - Sentieon DNAscope LongRead – A highly Accurate, Fast, and Efficient Pipeline for Germline Variant Calling from PacBio HiFi reads
- [UMI - BioRxiv 2022] - Processing UMI Datasets at High Accuracy and Efficiency with the Sentieon ctDNA Analysis Pipeline
- [TNscope - BioRxiv 2018] - TNscope: Accurate Detection of Somatic Mutations with Haplotype-based Variant Candidate Detection and Machine Learning Filtering
- [DNAseq/TNseq - BioRxiv 2017] - The Sentieon Genomics Tools - A fast and accurate solution to variant calling from next-generation sequence data
- [DNAseq - PeerJ 2016] - Sentieon DNA Pipeline for Variant Detection, Software-only solution, over 20X faster than GATK 3.3 with identical results
Selected Third Party Benchmark Publications:
- [DNAseq - Scientific Reports 2025] - Benchmarking of variant calling software for whole-exome sequencing using gold standard datasets
- [DNAscope - Research Square 2024] - Toward best practices for detecting germline small variants: a large-scale real-world WES benchmarking study using the Quartet DNA reference materials
- [DNAseq - MedRxiv 2024] - Rapid NGS Analysis on Google Cloud Platform: performance benchmark and user tutorial
- [DNAscope - BioRxiv 2022] - Sequencing by avidity enables high accuracy with low reagent consumption
- [DNAscope - BMC Genomics 2022] - Accuracy benchmark of the GeneMind GenoLab M sequencing platform for WGS and WES analysis
- [TNscope - Briefings in Bioinformatics 2020] - Benchmarking variant callers in next-generation and third-generation sequencing analysis
- [DNAscope - BioRxiv 2020] - Advanced Whole Genome Sequencing Using an Entirely PCR-free Massively Parallel Sequencing Workflow
- [DNAseq - Frontiers 2019] - Sentieon DNASeq Variant Calling Workflow Demonstrates Strong Computational Performance and Accuracy
Blogs and Use Cases
Selected blogs and use cases from industry collaborators
- Dell Technologies Whitepaper - Dell PowerEdge Servers Accelerate Genomics Data Processing with Sentieon Software on 5th Generation AMD EPYC Processors
- AWS for Industries - Enhanced Genomic Data Storage and Workflows with MGI, Sentieon, and AWS
- OmniTier Partner Solution Application Note - Benchmarking End-to-End, OnPremise, WGS Analysis, using Sentieon DNAscope Secondary Analysis with OmniTier Insight Tertiary Analysis
- Intel Health Analytics Solution Brief - Streamlining Genomic Analysis with Faster and More Accurate Technology
- Intel Developers Blog - Delivering Cost-Effective Genomics for Precision Medicine; Comparing Sentieon DNASeq Performance to NVIDIA Clara Parabricks
- Intel Health Analytics Solution Brief - Intel CPUs deliver breakthrough speed for genomic analysis
- Amazon Omics Blog - Amazon Omics now supports Sentieon genomic analysis pipelines
- Microsoft Healthcare and Life Sciences Blog - Sentieon pipelines on Azure – An overview
- Oracle Cloud Infrastructure Blog - Oracle Cloud and Sentieon DNAseq enable processing of 30X WGS samples from FASTQ to VCF in about an hour and less than $1 in on-demand costs
- ByteDance Volcengine Blog - (written in Chinese) Volcano Engine Bio-OS Platform empowers Sentieon to achieve efficient and precise genomic analysis
- Golden Helix Blog - Sentieon’s Latest Enhancements Powering VarSeq 2.5.0: Accelerating Clinical Workflows and Data Analysis in NGS
- Golden Helix Blog - Top 5 Features of Sentieon
- Golden Helix Blog - Sentieon Updates: Now Supporting Long-Read Alignment and Calling
- DNAnexus Science Frontiers - An exploration of machine-learning based variant callers for SNV and small-indel using PacBio HiFi data
- DNANexus Case Study - Calling Somatic Variants with Sentieon on DNANexus
- DNANexus Case Study - Trio Analysis with Sentieon Rapid DNASeq on DNAnexus
- Memory Machine Cloud Blog - Sentieon Genomics Tools: Boosting Performance with Memory Machine Cloud (complete report)