DNAscope LongRead Nanopore Pipeline

- Optimized pipeline for SNP/Indel/SV detection;
- Higher accuracy than Clair3 and Sniffles2;
- DNAscope for ONT is 3-5x faster;
Access to current DNAscope ONT Performance Datasheet (PDF version)
Fast and Accurate SNP/Indel Calling Pipeline
- Integrated pipeline from fastq to VCF, perform alignment and variant calling;
- Output GVCF and VCF files; Supports Chemistry R10.4 and above;
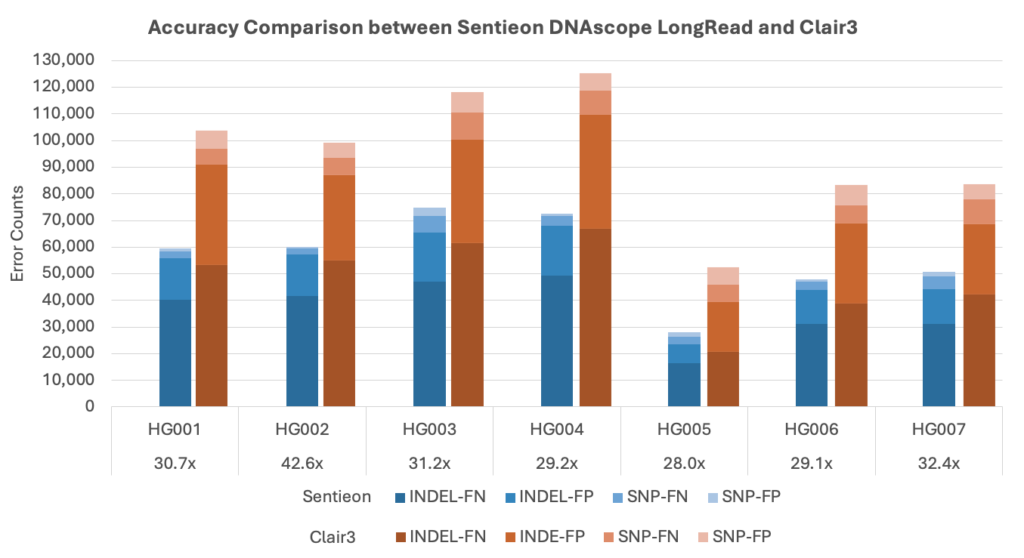
- Compared to Clair3, Sentieon pipeline reduces SNP and Indel error by over 50%;
- Preprint published on BioRxiv;
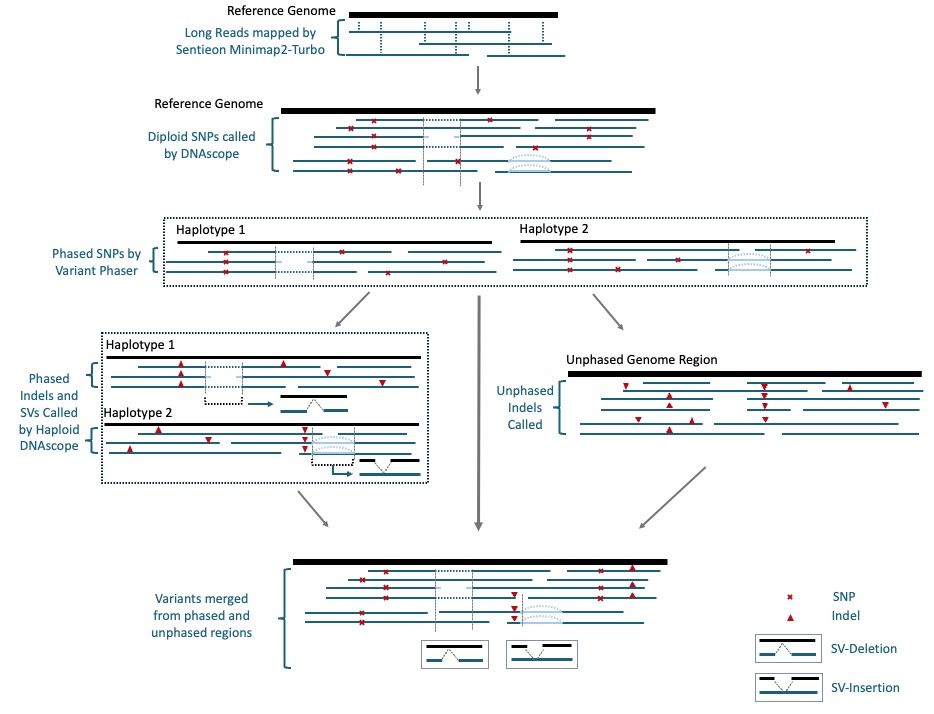
Schematic Diagram of DNAscope LongRead Pipeline: Part of the Genome was assembled into haplotypes to generate more accurate Indel and SV variants.
More than 50% SNP error reduction compared to Clair3.
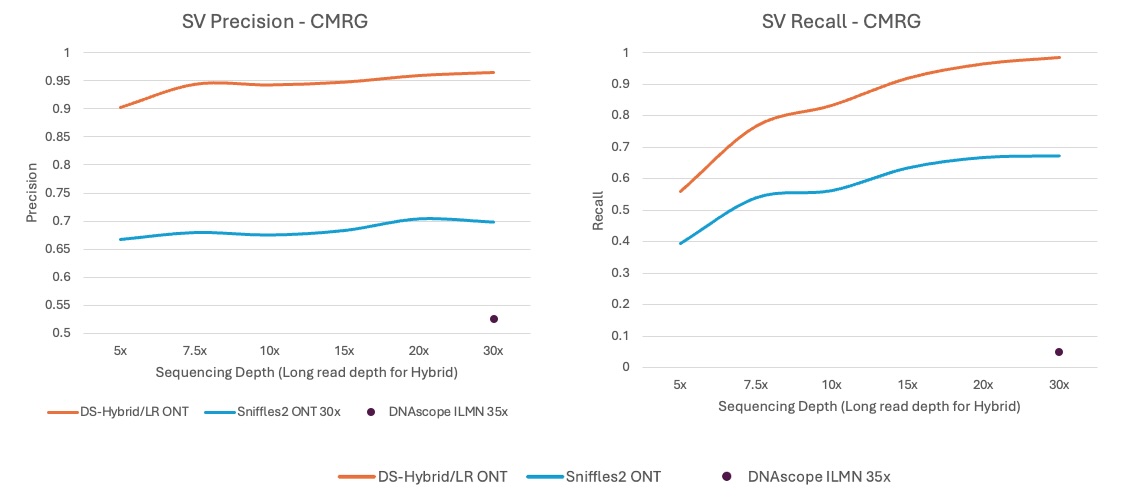
Highly Accurate Structural Variants Calling Pipeline
Haplotype-resolved single base resolution SV caller, higher accuracy than Sniffles2 on CMRG regions.
Rapid Alignment and Variant Calling
- Complete CRAM-VCF analysis within as fast as 13min.
- Peak Memory under 35GB.
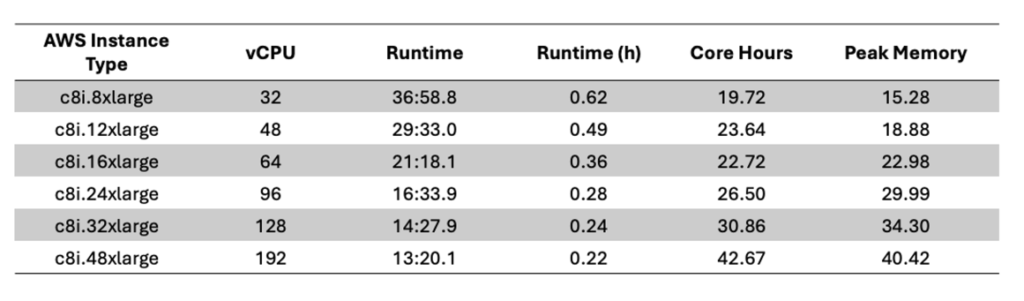
How fast and how cheap? Efficiency metrics for DNAscope LongRead on ONT WGS data (from CRAM to VCF).